Tissue and Cell Biophysics
Description
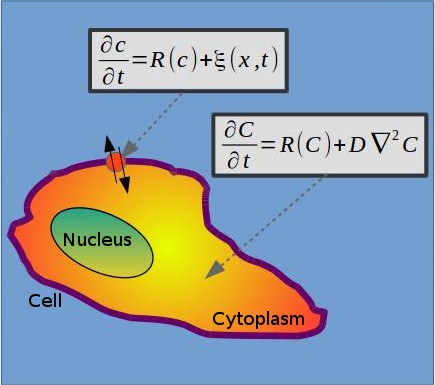
- Understanding of experimental results on cell motion obtained with amoebae cells. Adaptation of complex biochemical models, together with a phase field approach, to study specific biochemical characteristics of the experimental profiles of the different biochemical markers of intracellular waves. Study of confinement of cells and interactions with borders or objects to evaluate the carrying properties observed in certain cells. Methods: SPDE with reaction-diffusion equations and Ornstein-Uhlenbeck processes.
- Stochastic dynamics of ion channels, both individually and coupled to other channels. Dynamics subconductance states of v-gated channels and dynamics of coupled K, Na, and Ca channels. Firing bursts in the study of patterns of Purkinje neurons. Methods: Langevin equations with non-trivial boundary conditions (coupling of thee model different types of biological processes occurring in the interior of living cells using techniques previously developed in physics and mathematics. We consider problems at different scales, from the stochastic behaviour of individual ion gates to the spatial particularities of coupled opposite enzymatic reactions or the viscoelastic properties of the cytoskeleton of cells. Analytic calculations together with the use of numerical simulations permit to reveal some of the mechanisms involved in the biological processes inside of the cells and their relation with the external responses to small perturbations like cell excitations of neurons or cell locomotion.
- Self-organization of embryonic organoid, Reconstitution studies using embryonic stem cells have revealed that 3D aggregates of cells are able to spontaneously polarize in vitro and elongate, mimicking the early formation of body axes during animal development. One such example are gastruloids, embryonic stem cell aggregates that recapitulate the axial organization of post-implantation embryos. Gastruloids currently stand as a promising system for personalized medicine and drug testing. By combining soft matter approaches, agent-based modelling and image-based experimental data, we aim to elucidate the biophysical basis of spontaneous symmetry breaking and elongation of such structures. Methods: stochastic processes, systems biology, gene regulatory networks, agent-based modelling, soft matter physics, image segmentation and data analysis.
Contact: Laureano Ramirez, Sergio Alonso , David Oriola
Share: